peptides
spectra
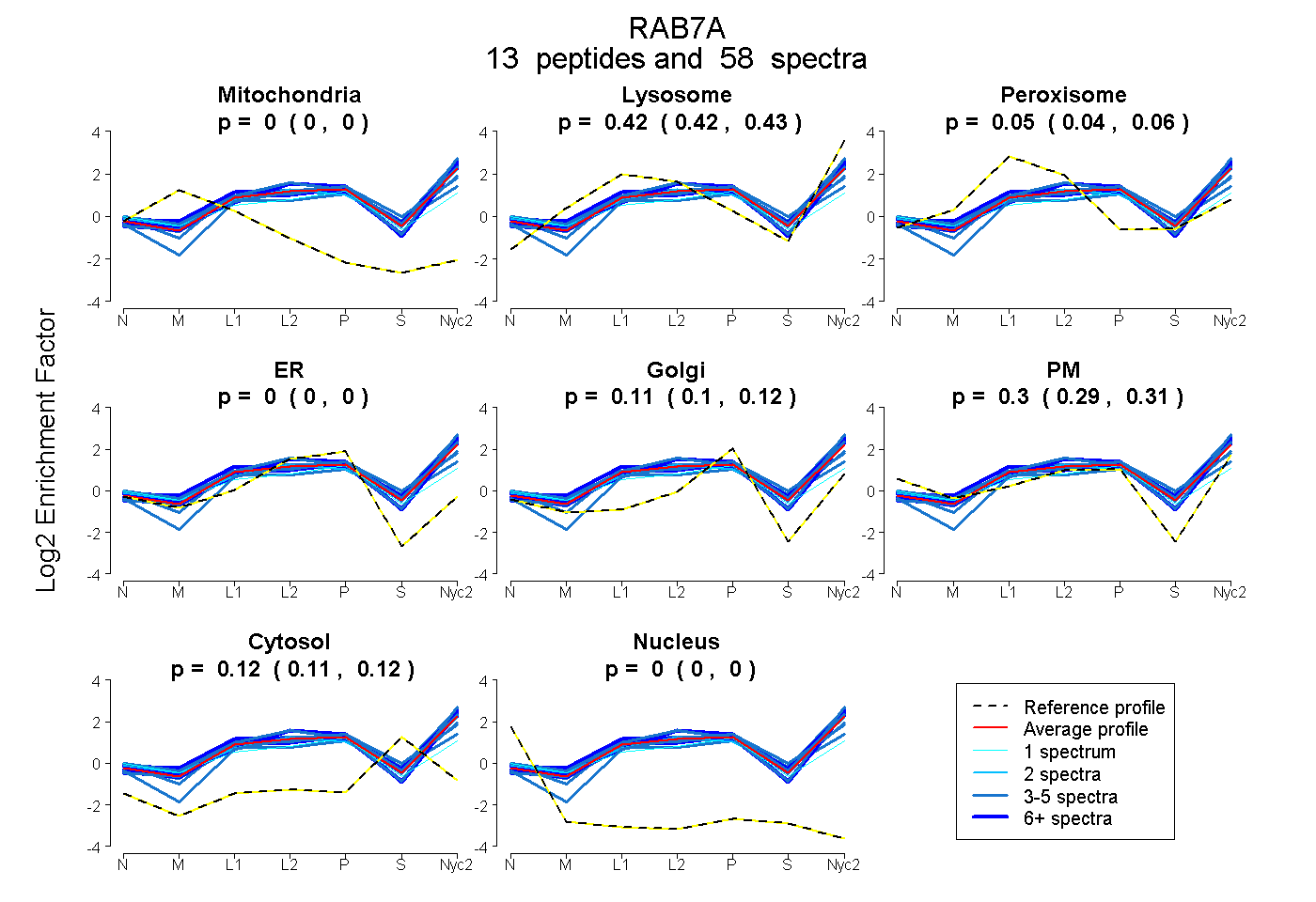
0.000 | 0.000
0.415 | 0.432
0.038 | 0.058
0.000 | 0.000
0.102 | 0.121
0.287 | 0.308
0.112 | 0.118
0.000 | 0.000
peptides
spectra

0.000 | 0.000
0.592 | 0.602
0.000 | 0.000
0.344 | 0.357
0.000 | 0.000
0.045 | 0.056
0.000 | 0.000
| Plot | Mito | Lyso | Perox | ER | Golgi | PM | Cytosol | Nucleus | |||||
| Expt A |
peptides |
58 spectra |
 |
0.000 0.000 | 0.000 |
0.425 0.415 | 0.432 |
0.049 0.038 | 0.058 |
0.000 0.000 | 0.000 |
0.112 0.102 | 0.121 |
0.299 0.287 | 0.308 |
0.115 0.112 | 0.118 |
0.000 0.000 | 0.000 |
||
| Plot | Mito | Lyso or Perox | ER | Golgi | PM | Cytosol | Nucleus | ||||||
| Expt B |
peptides |
29 spectra |
 |
0.000 0.000 | 0.000 |
0.598 0.592 | 0.602 |
0.000 0.000 | 0.000 |
0.351 0.344 | 0.357 |
0.000 0.000 | 0.000 |
0.051 0.045 | 0.056 |
0.000 0.000 | 0.000 |
| 2 spectra, EVMVDDR | 0.000 | 0.455 | 0.328 | 0.000 | 0.000 | 0.217 | 0.000 | |||
| 9 spectra, DPENFPFVVLGNK | 0.000 | 0.578 | 0.024 | 0.398 | 0.000 | 0.000 | 0.000 | |||
| 2 spectra, FSNQYK | 0.000 | 0.544 | 0.128 | 0.268 | 0.000 | 0.060 | 0.000 | |||
| 4 spectra, EAINVEQAFQTIAR | 0.001 | 0.579 | 0.000 | 0.350 | 0.000 | 0.069 | 0.000 | |||
| 2 spectra, ATIGADFLTK | 0.000 | 0.550 | 0.168 | 0.282 | 0.000 | 0.000 | 0.000 | |||
| 1 spectrum, DEFLIQASPR | 0.000 | 0.472 | 0.000 | 0.324 | 0.000 | 0.204 | 0.000 | |||
| 2 spectra, NNIPYFETSAK | 0.000 | 0.490 | 0.000 | 0.257 | 0.184 | 0.069 | 0.000 | |||
| 3 spectra, FQSLGVAFYR | 0.000 | 0.696 | 0.000 | 0.300 | 0.000 | 0.004 | 0.000 | |||
| 1 spectrum, GADCCVLVFDVTAPNTFK | 0.000 | 0.717 | 0.028 | 0.250 | 0.000 | 0.005 | 0.000 | |||
| 3 spectra, VIILGDSGVGK | 0.000 | 0.565 | 0.019 | 0.295 | 0.000 | 0.121 | 0.000 |
| Plot | Lyso | Other | |||||||||||
| Expt C |
peptides |
185 spectra |
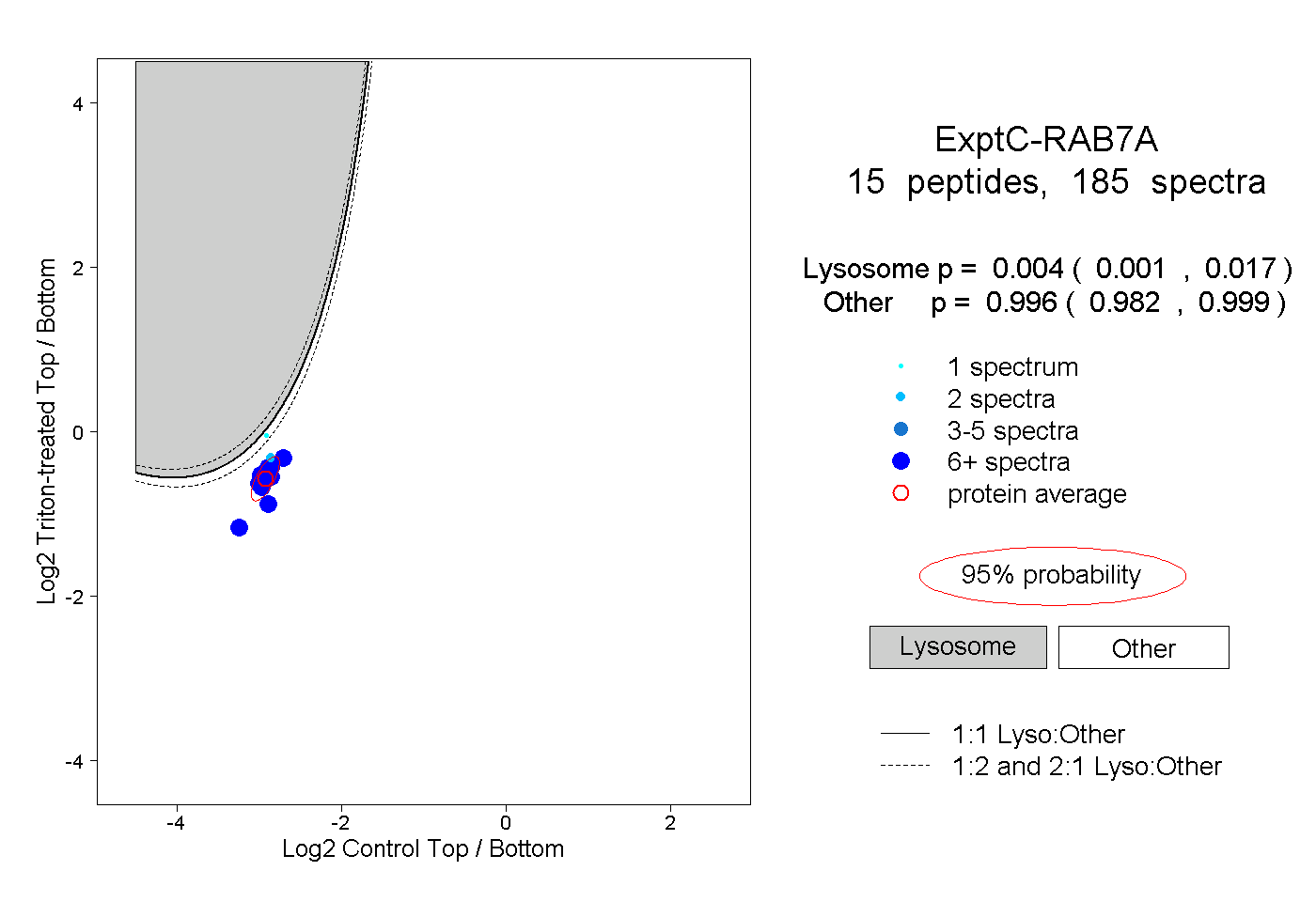 |
0.004 0.001 | 0.017 |
0.996 0.982 | 0.999 |
||||||||
| Plot | Lyso | Other | |||||||||||
| Expt D |
peptides |
10 spectra |
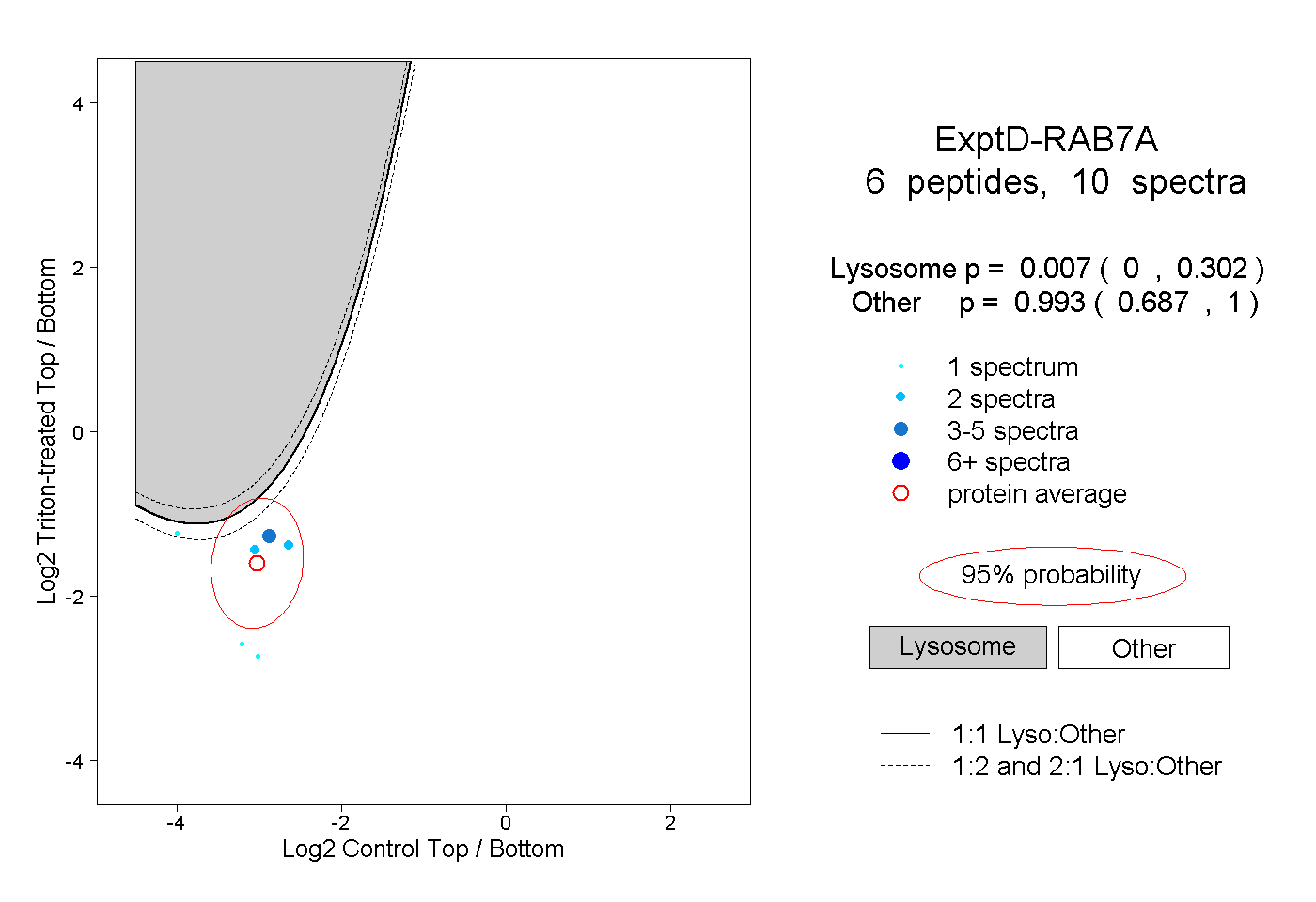 |
0.007 0.000 | 0.302 |
0.993 0.687 | 1.000 |