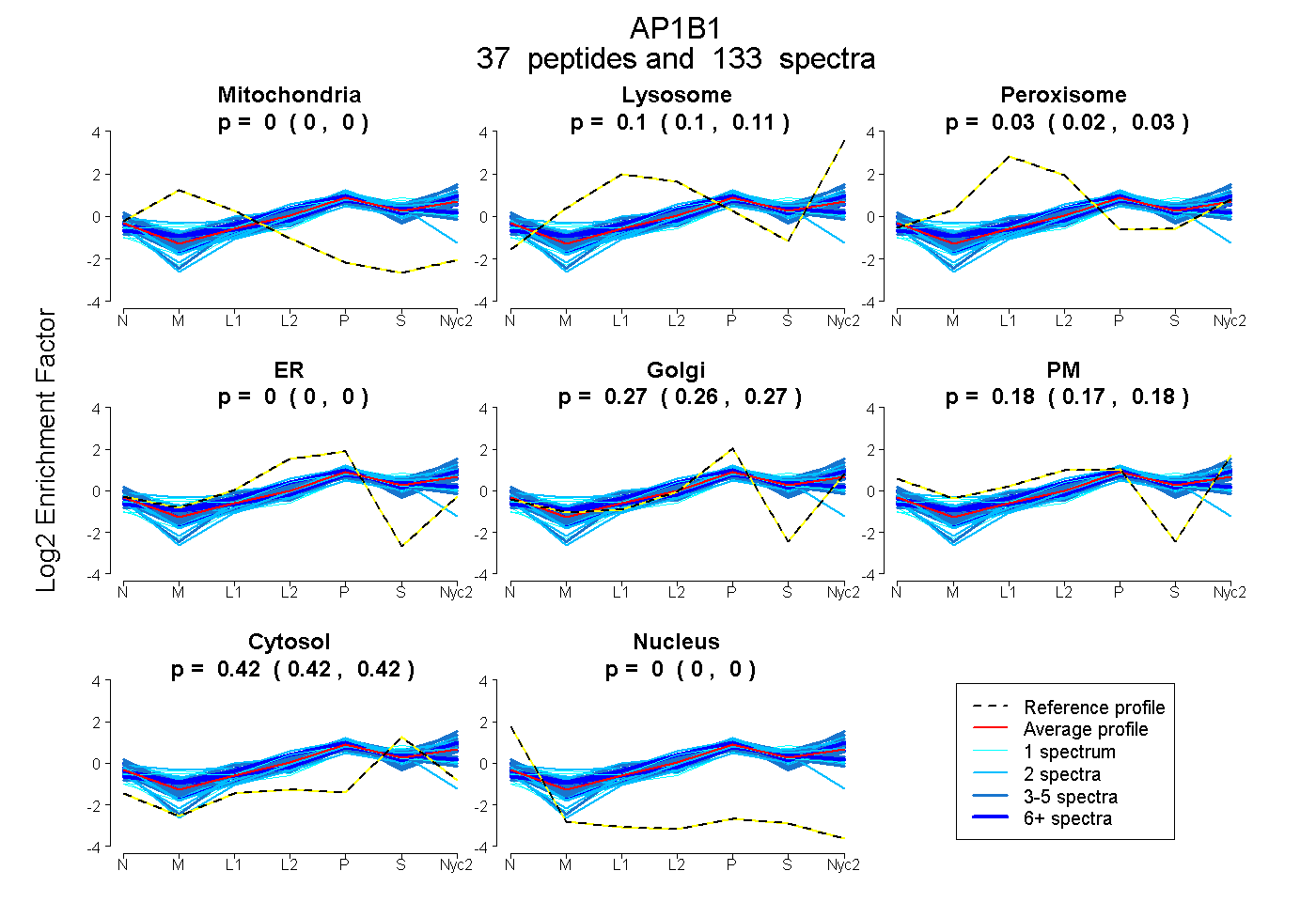
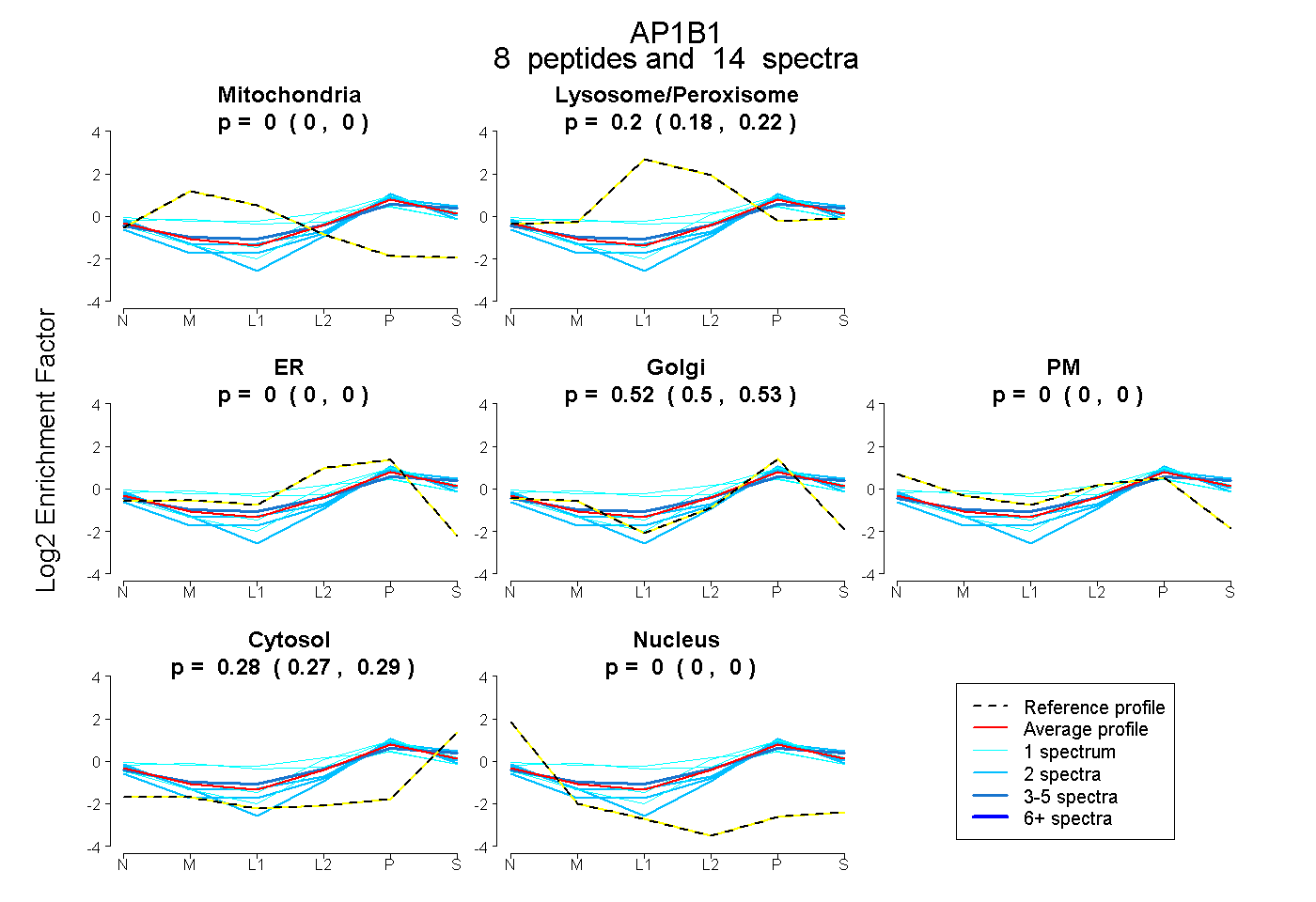
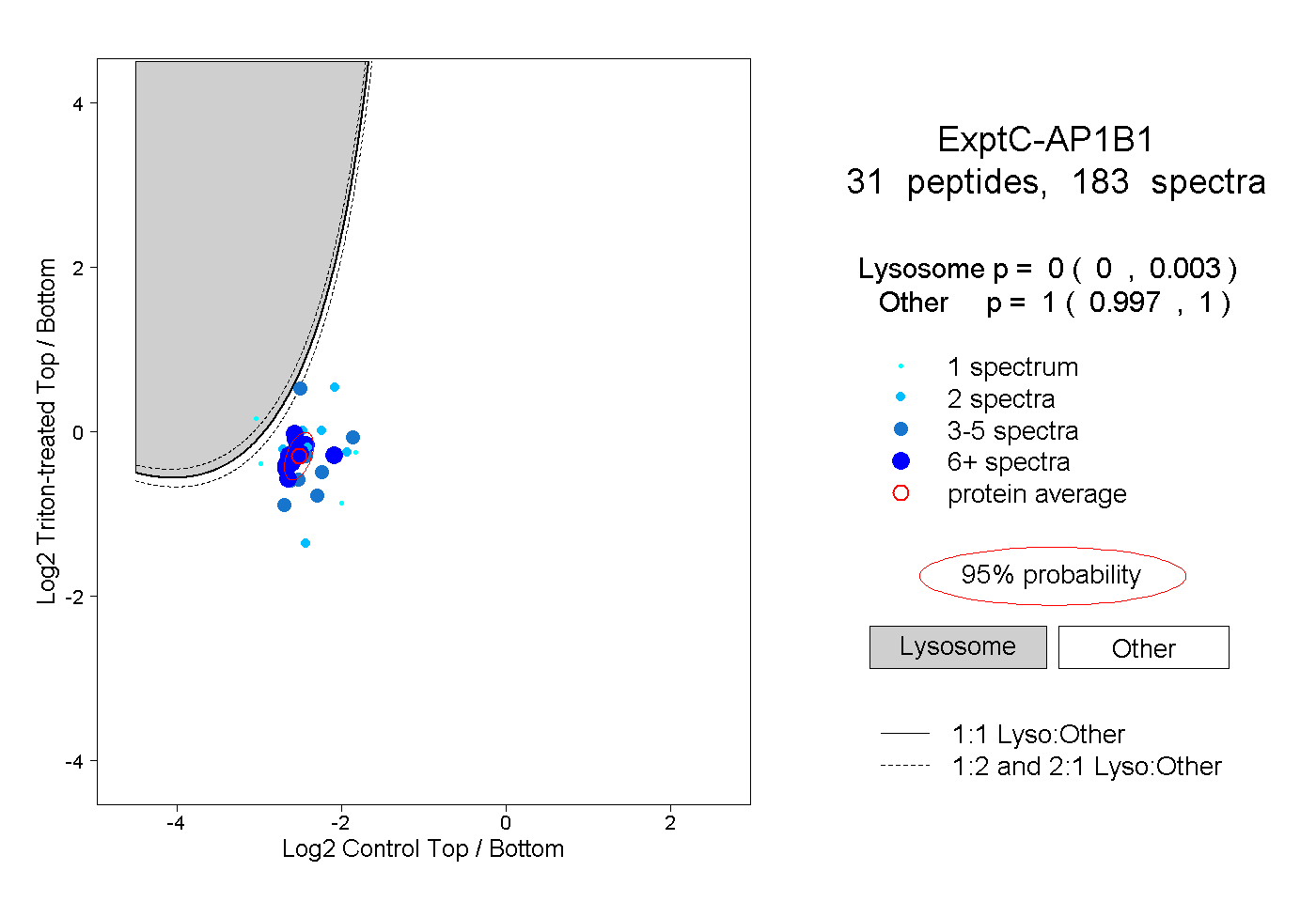
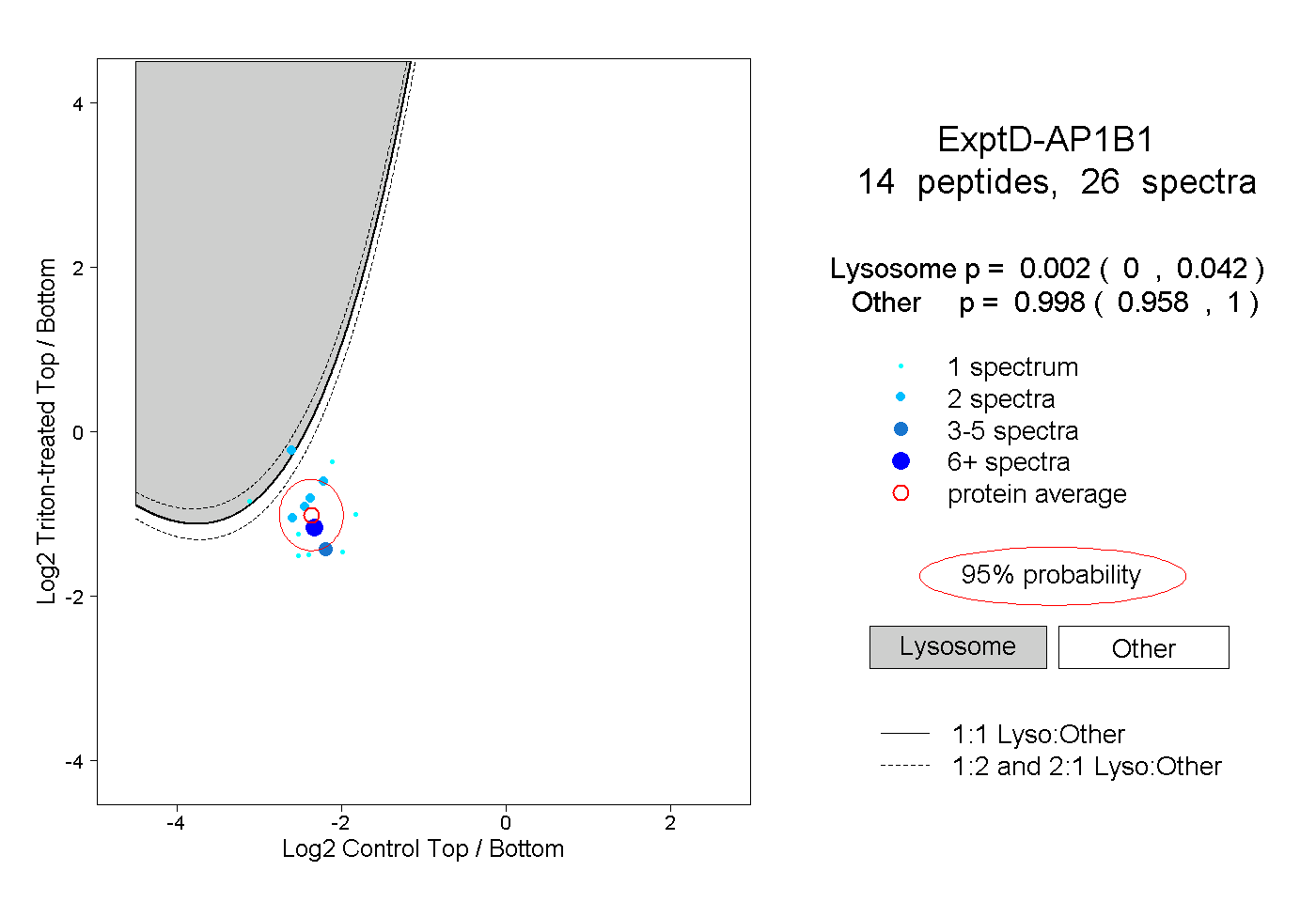
| 9 spectra, YFTTTK |
|
0.000 |
|
|
|
|
|
|
|
1.000 |
| 4 spectra, TAAVCVAK |
|
0.000 |
|
|
|
|
|
|
|
1.000 |
| 11 spectra, GYIYWR |
|
0.001 |
|
|
|
|
|
|
|
0.999 |
| 4 spectra, FMEMLSK |
|
0.000 |
|
|
|
|
|
|
|
1.000 |
| 1 spectrum, LSHANSAVVLSAVK |
|
0.039 |
|
|
|
|
|
|
|
0.961 |
| 2 spectra, APEVSQHVYQAYETILK |
|
0.000 |
|
|
|
|
|
|
|
1.000 |
| 8 spectra, LLSTDPVAAK |
|
0.008 |
|
|
|
|
|
|
|
0.992 |
| 1 spectrum, LASQANIAQVLAELK |
|
0.000 |
|
|
|
|
|
|
|
1.000 |
| 1 spectrum, LAPPLVTLLSAEPELQYVALR |
|
0.930 |
|
|
|
|
|
|
|
0.070 |
| 2 spectra, GEIFELK |
|
0.009 |
|
|
|
|
|
|
|
0.991 |
| 8 spectra, DLDYYATLLK |
|
0.000 |
|
|
|
|
|
|
|
1.000 |
| 21 spectra, RPEILK |
|
0.000 |
|
|
|
|
|
|
|
1.000 |
| 7 spectra, QMFLATWK |
|
0.000 |
|
|
|
|
|
|
|
1.000 |
| 2 spectra, IQPGNPSFTLSLK |
|
0.000 |
|
|
|
|
|
|
|
1.000 |
| 2 spectra, AELNSDK |
|
0.008 |
|
|
|
|
|
|
|
0.992 |
| 2 spectra, LHDINAQLVEDQGFLDTLK |
|
0.000 |
|
|
|
|
|
|
|
1.000 |
| 5 spectra, AVWLPAMK |
|
0.000 |
|
|
|
|
|
|
|
1.000 |
| 3 spectra, LQSSNIFTVAK |
|
0.000 |
|
|
|
|
|
|
|
1.000 |
| 11 spectra, DCEDPNPLIR |
|
0.001 |
|
|
|
|
|
|
|
0.999 |
| 4 spectra, DEDPYVR |
|
0.161 |
|
|
|
|
|
|
|
0.839 |
| 6 spectra, YNDPIYVK |
|
0.001 |
|
|
|
|
|
|
|
0.999 |
| 2 spectra, MEPLNNLQVAVK |
|
0.000 |
|
|
|
|
|
|
|
1.000 |
| 2 spectra, VNYVVQEAIVVIK |
|
0.000 |
|
|
|
|
|
|
|
1.000 |
| 28 spectra, VIASMTVGK |
|
0.003 |
|
|
|
|
|
|
|
0.997 |
| 10 spectra, NINLIVQK |
|
0.001 |
|
|
|
|
|
|
|
0.999 |
| 3 spectra, AAMIWIVGEYAER |
|
0.000 |
|
|
|
|
|
|
|
1.000 |
| 1 spectrum, DIPNENEAQFQIR |
|
0.000 |
|
|
|
|
|
|
|
1.000 |
| 2 spectra, CVSTLLDLIQTK |
|
0.002 |
|
|
|
|
|
|
|
0.998 |
| 4 spectra, ITEYLCEPLR |
|
0.000 |
|
|
|
|
|
|
|
1.000 |
| 4 spectra, DCPLNTEAASSK |
|
0.000 |
|
|
|
|
|
|
|
1.000 |
| 13 spectra, LDIMIR |
|
0.002 |
|
|
|
|
|
|
|
0.998 |