peptides
spectra
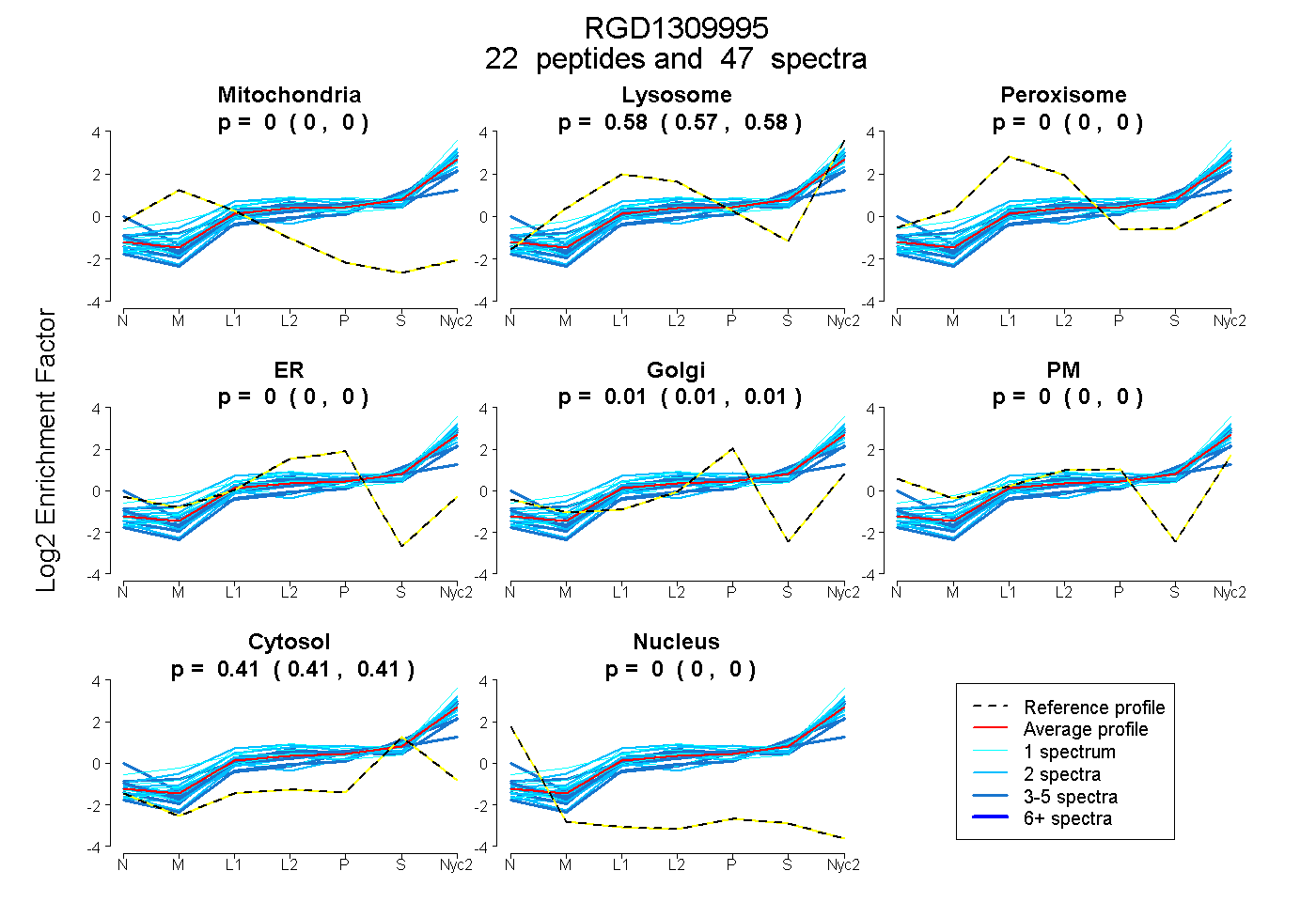
0.000 | 0.000
0.572 | 0.581
0.000 | 0.000
0.000 | 0.000
0.007 | 0.015
0.000 | 0.000
0.408 | 0.414
0.000 | 0.000
peptides
spectra
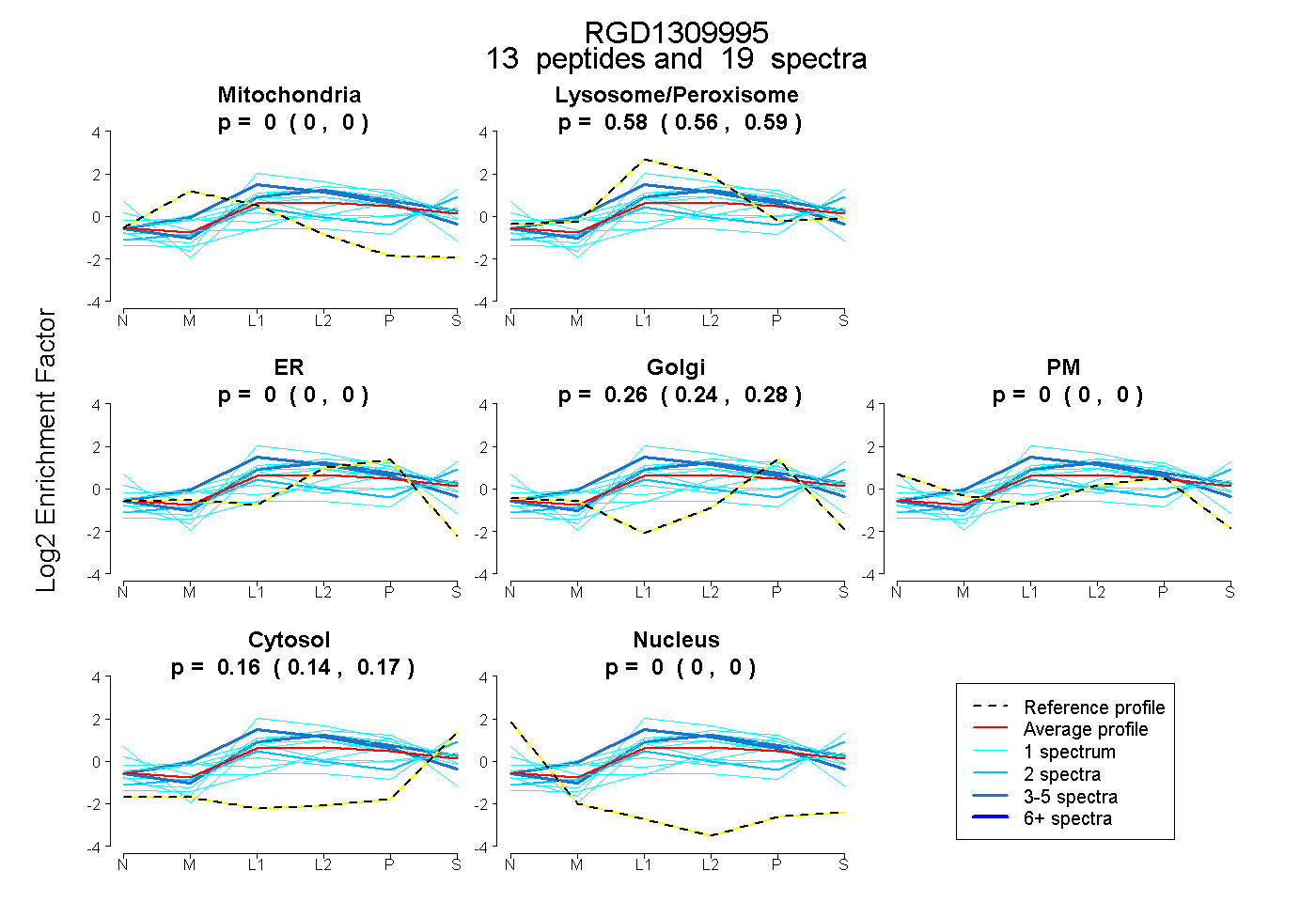
0.000 | 0.000
0.563 | 0.592
0.000 | 0.000
0.241 | 0.278
0.000 | 0.000
0.143 | 0.172
0.000 | 0.000
| Plot | Mito | Lyso | Perox | ER | Golgi | PM | Cytosol | Nucleus | |||||
| Expt A |
peptides |
47 spectra |
 |
0.000 0.000 | 0.000 |
0.577 0.572 | 0.581 |
0.000 0.000 | 0.000 |
0.000 0.000 | 0.000 |
0.012 0.007 | 0.015 |
0.000 0.000 | 0.000 |
0.412 0.408 | 0.414 |
0.000 0.000 | 0.000 |
||
| Plot | Mito | Lyso or Perox | ER | Golgi | PM | Cytosol | Nucleus | ||||||
| Expt B |
peptides |
19 spectra |
 |
0.000 0.000 | 0.000 |
0.579 0.563 | 0.592 |
0.000 0.000 | 0.000 |
0.263 0.241 | 0.278 |
0.000 0.000 | 0.000 |
0.159 0.143 | 0.172 |
0.000 0.000 | 0.000 |
| 1 spectrum, FGIQEER | 0.051 | 0.529 | 0.000 | 0.358 | 0.000 | 0.062 | 0.000 | |||
| 4 spectra, LGEPSEIDQR | 0.000 | 0.583 | 0.234 | 0.034 | 0.000 | 0.148 | 0.000 | |||
| 1 spectrum, QLLSQIGNAMGYIR | 0.000 | 0.342 | 0.000 | 0.076 | 0.296 | 0.286 | 0.000 | |||
| 1 spectrum, MLVDVFAPEFR | 0.000 | 0.224 | 0.091 | 0.369 | 0.316 | 0.000 | 0.000 | |||
| 1 spectrum, LLPYEQSSLLELIK | 0.000 | 0.519 | 0.388 | 0.000 | 0.000 | 0.093 | 0.000 | |||
| 1 spectrum, FLEEYTSQLR | 0.000 | 0.595 | 0.269 | 0.021 | 0.000 | 0.116 | 0.000 | |||
| 1 spectrum, EEGLAEETLR | 0.000 | 0.548 | 0.000 | 0.029 | 0.299 | 0.070 | 0.055 | |||
| 1 spectrum, YLYNLNNQIFIER | 0.003 | 0.468 | 0.000 | 0.242 | 0.000 | 0.286 | 0.000 | |||
| 1 spectrum, LLQTMNLTQK | 0.000 | 0.614 | 0.000 | 0.273 | 0.000 | 0.113 | 0.000 | |||
| 1 spectrum, LLEILK | 0.000 | 0.766 | 0.184 | 0.000 | 0.000 | 0.051 | 0.000 | |||
| 3 spectra, NIHIFVSR | 0.000 | 0.733 | 0.000 | 0.267 | 0.000 | 0.000 | 0.000 | |||
| 2 spectra, DAFLQQK | 0.000 | 0.529 | 0.000 | 0.017 | 0.000 | 0.454 | 0.000 | |||
| 1 spectrum, AYVTHYLDK | 0.000 | 0.334 | 0.000 | 0.000 | 0.000 | 0.666 | 0.000 |
| Plot | Lyso | Other | |||||||||||
| Expt C |
peptides |
147 spectra |
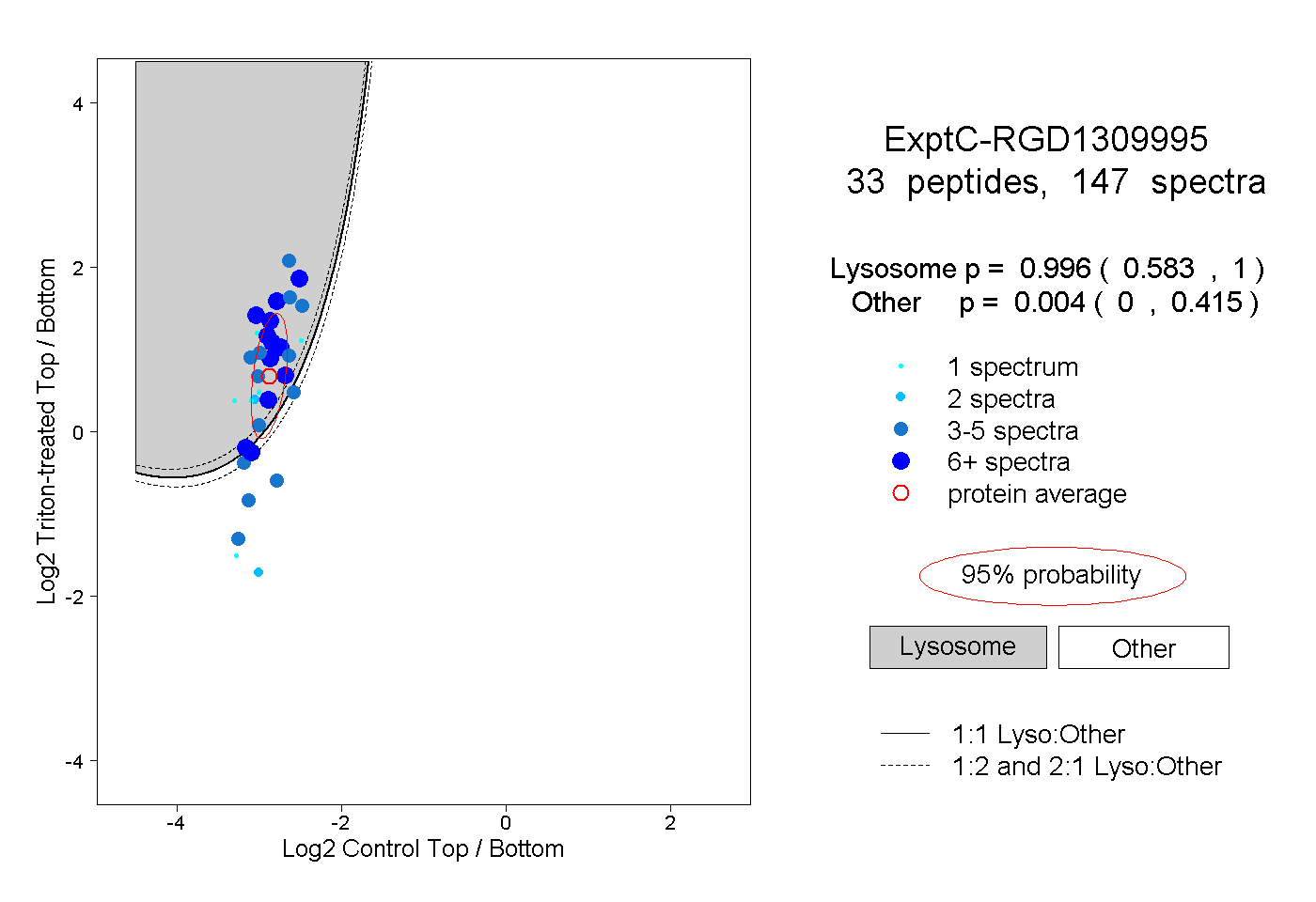 |
0.996 0.583 | 1.000 |
0.004 0.000 | 0.415 |
||||||||
| Plot | Lyso | Other | |||||||||||
| Expt D |
peptides |
21 spectra |
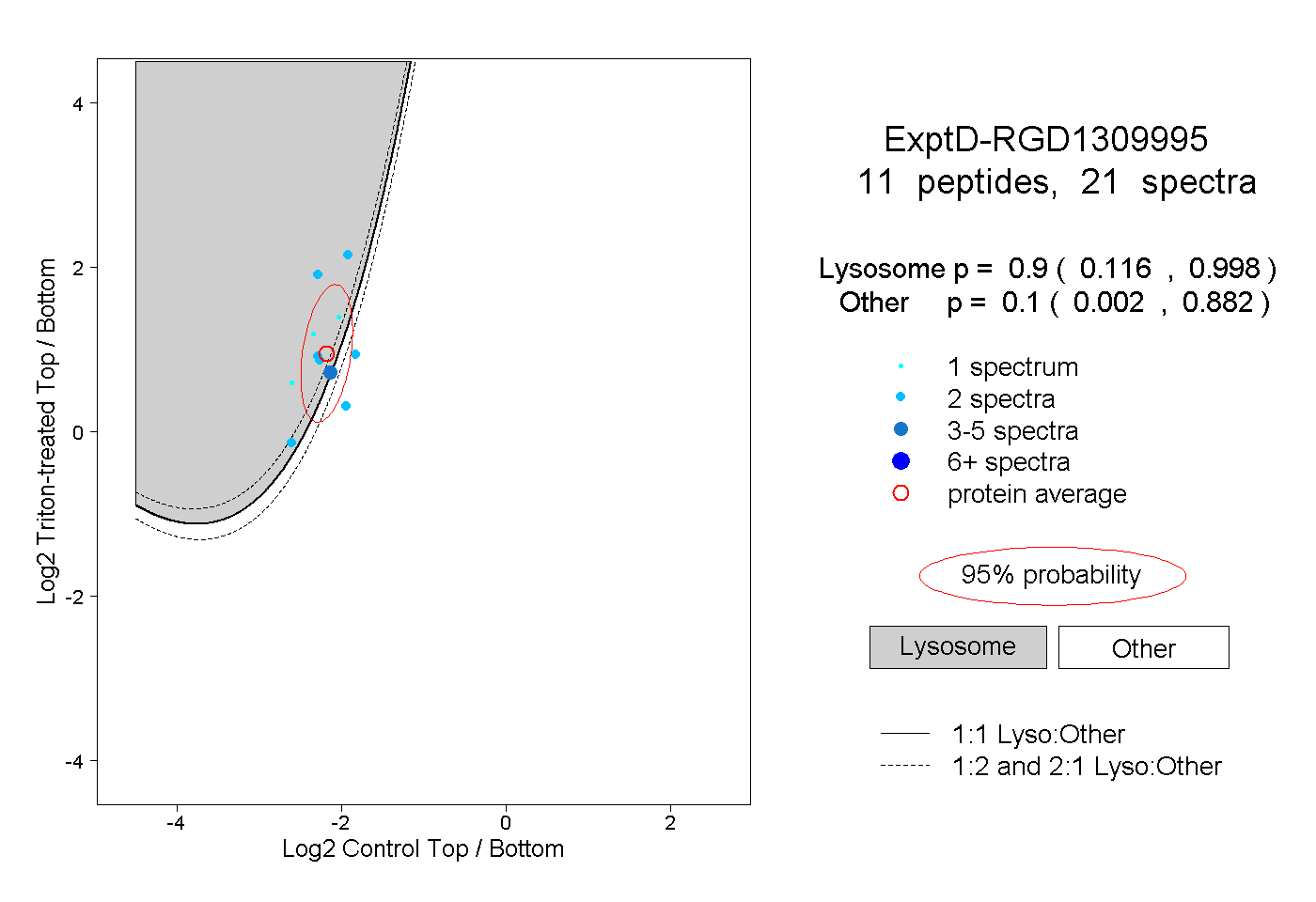 |
0.900 0.116 | 0.998 |
0.100 0.002 | 0.882 |