peptides
spectra
0.000 | 0.000
0.374 | 0.381
0.000 | 0.000
0.000 | 0.000
0.076 | 0.082
0.000 | 0.000
0.540 | 0.545
0.000 | 0.000
peptides
spectra
0.000 | 0.000
0.439 | 0.450
0.000 | 0.000
0.115 | 0.132
0.000 | 0.000
0.422 | 0.438
0.000 | 0.000
| Plot | Mito | Lyso | Perox | ER | Golgi | PM | Cytosol | Nucleus | |||||
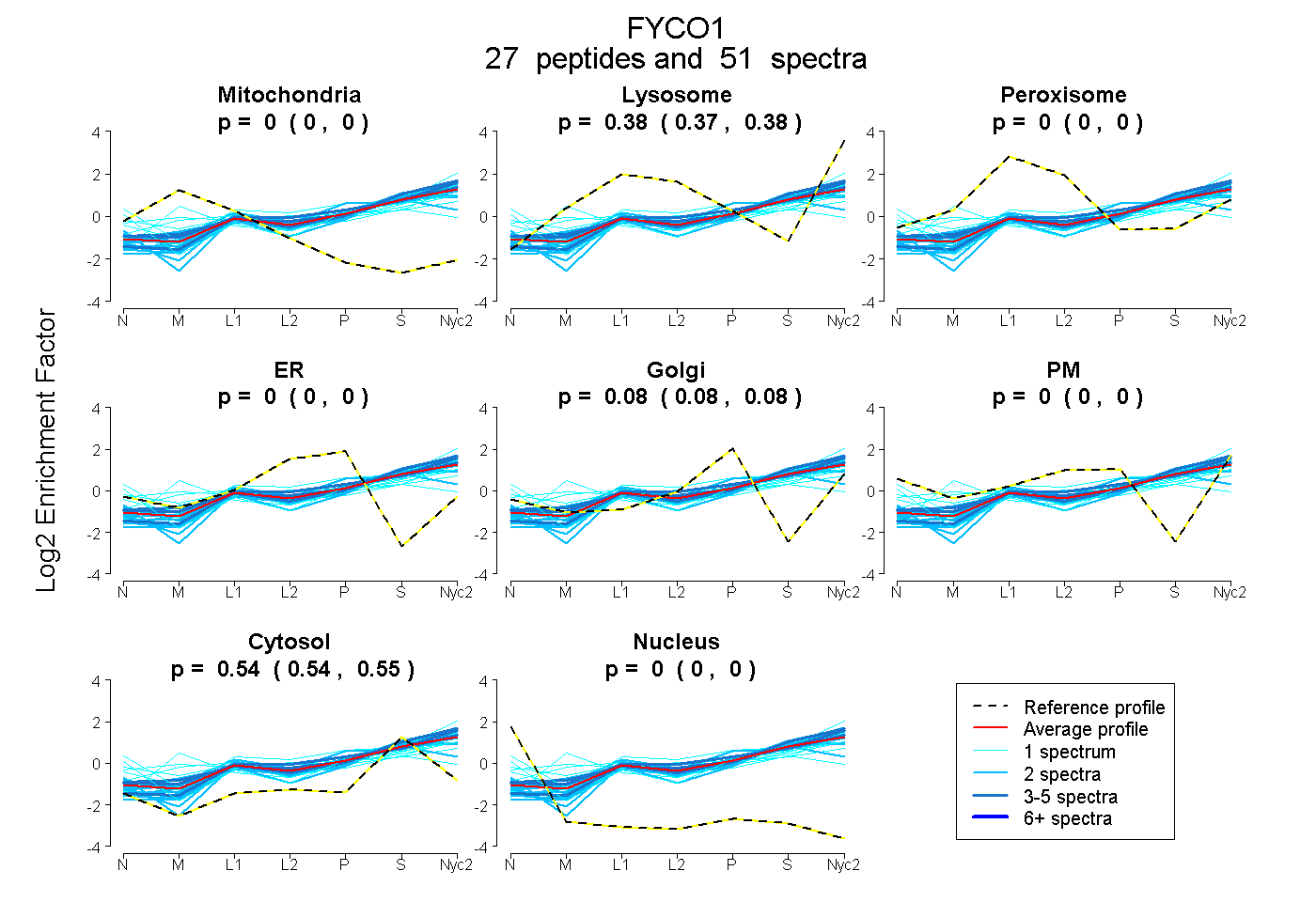
| Expt A |
peptides |
51 spectra |
 |
0.000 0.000 | 0.000 |
0.378 0.374 | 0.381 |
0.000 0.000 | 0.000 |
0.000 0.000 | 0.000 |
0.079 0.076 | 0.082 |
0.000 0.000 | 0.000 |
0.543 0.540 | 0.545 |
0.000 0.000 | 0.000 |
||
| Plot | Mito | Lyso or Perox | ER | Golgi | PM | Cytosol | Nucleus | ||||||
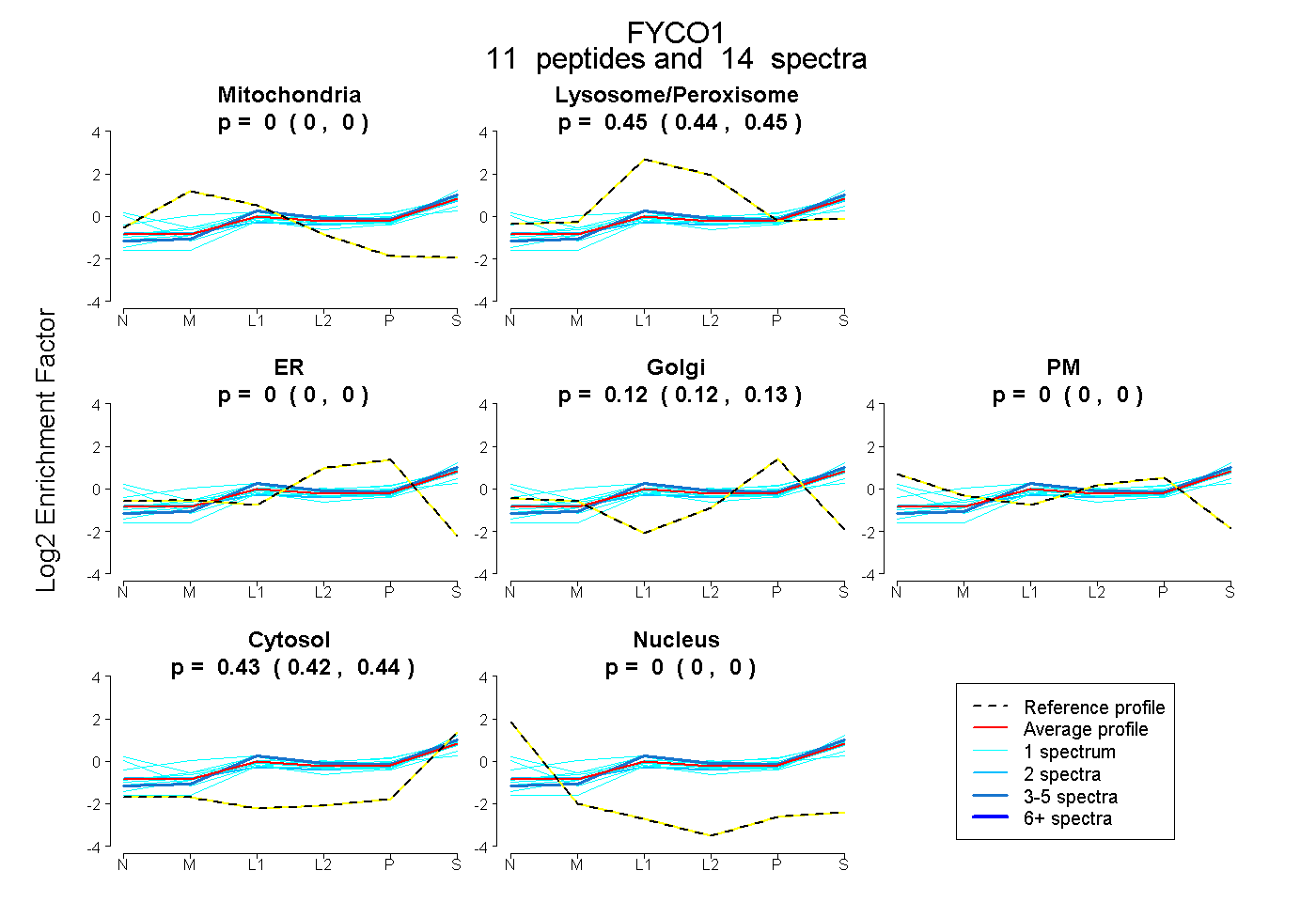
| Expt B |
peptides |
14 spectra |
 |
0.000 0.000 | 0.000 |
0.445 0.439 | 0.450 |
0.000 0.000 | 0.000 |
0.124 0.115 | 0.132 |
0.000 0.000 | 0.000 |
0.431 0.422 | 0.438 |
0.000 0.000 | 0.000 |
| 1 spectrum, EGILQEESIYK | 0.000 | 0.438 | 0.000 | 0.164 | 0.000 | 0.398 | 0.000 | |||
| 1 spectrum, LEYLLQFDQK | 0.000 | 0.414 | 0.024 | 0.000 | 0.000 | 0.562 | 0.000 | |||
| 1 spectrum, EAALQER | 0.000 | 0.332 | 0.000 | 0.000 | 0.243 | 0.425 | 0.000 | |||
| 1 spectrum, AHIQELLQCSER | 0.087 | 0.440 | 0.000 | 0.222 | 0.000 | 0.250 | 0.000 | |||
| 1 spectrum, QLQEHVQQLNR | 0.000 | 0.350 | 0.000 | 0.000 | 0.278 | 0.347 | 0.026 | |||
| 1 spectrum, ALQAELSQVR | 0.000 | 0.493 | 0.000 | 0.110 | 0.000 | 0.396 | 0.000 | |||
| 2 spectra, DAVPLQEELSGK | 0.000 | 0.392 | 0.000 | 0.159 | 0.000 | 0.449 | 0.000 | |||
| 1 spectrum, ALENQCQQQIQLIEVLSAEK | 0.000 | 0.448 | 0.023 | 0.098 | 0.000 | 0.431 | 0.000 | |||
| 3 spectra, VLIPTTR | 0.000 | 0.488 | 0.000 | 0.055 | 0.000 | 0.457 | 0.000 | |||
| 1 spectrum, LNCTGSHLAECQATLLR | 0.000 | 0.395 | 0.000 | 0.113 | 0.000 | 0.492 | 0.000 | |||
| 1 spectrum, LCQEVTNR | 0.000 | 0.469 | 0.000 | 0.091 | 0.000 | 0.440 | 0.000 |
| Plot | Lyso | Other | |||||||||||
| Expt C |
peptides |
84 spectra |
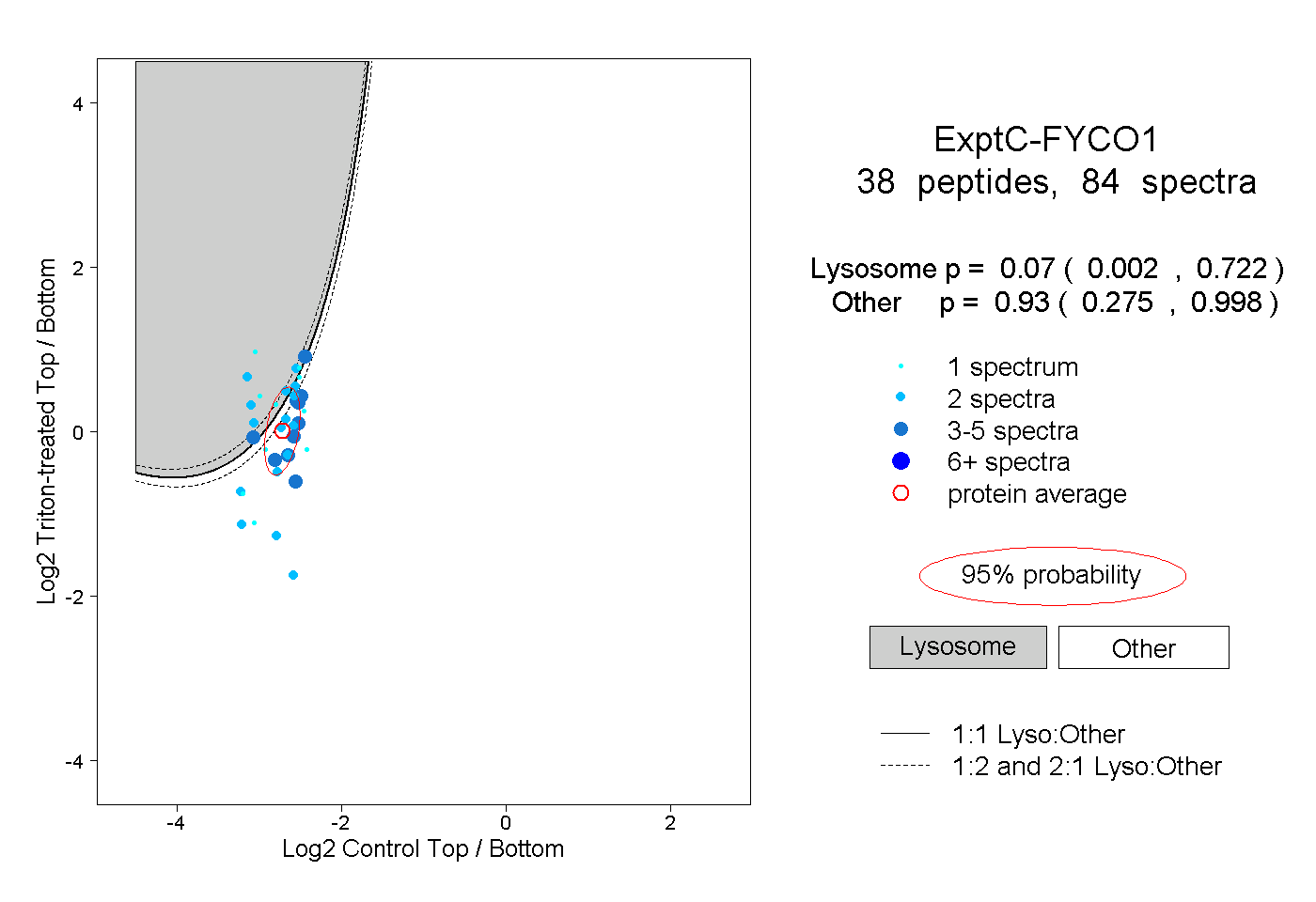 |
0.070 0.002 | 0.722 |
0.930 0.275 | 0.998 |
||||||||
| Plot | Lyso | Other | |||||||||||
| Expt D |
peptides |
14 spectra |
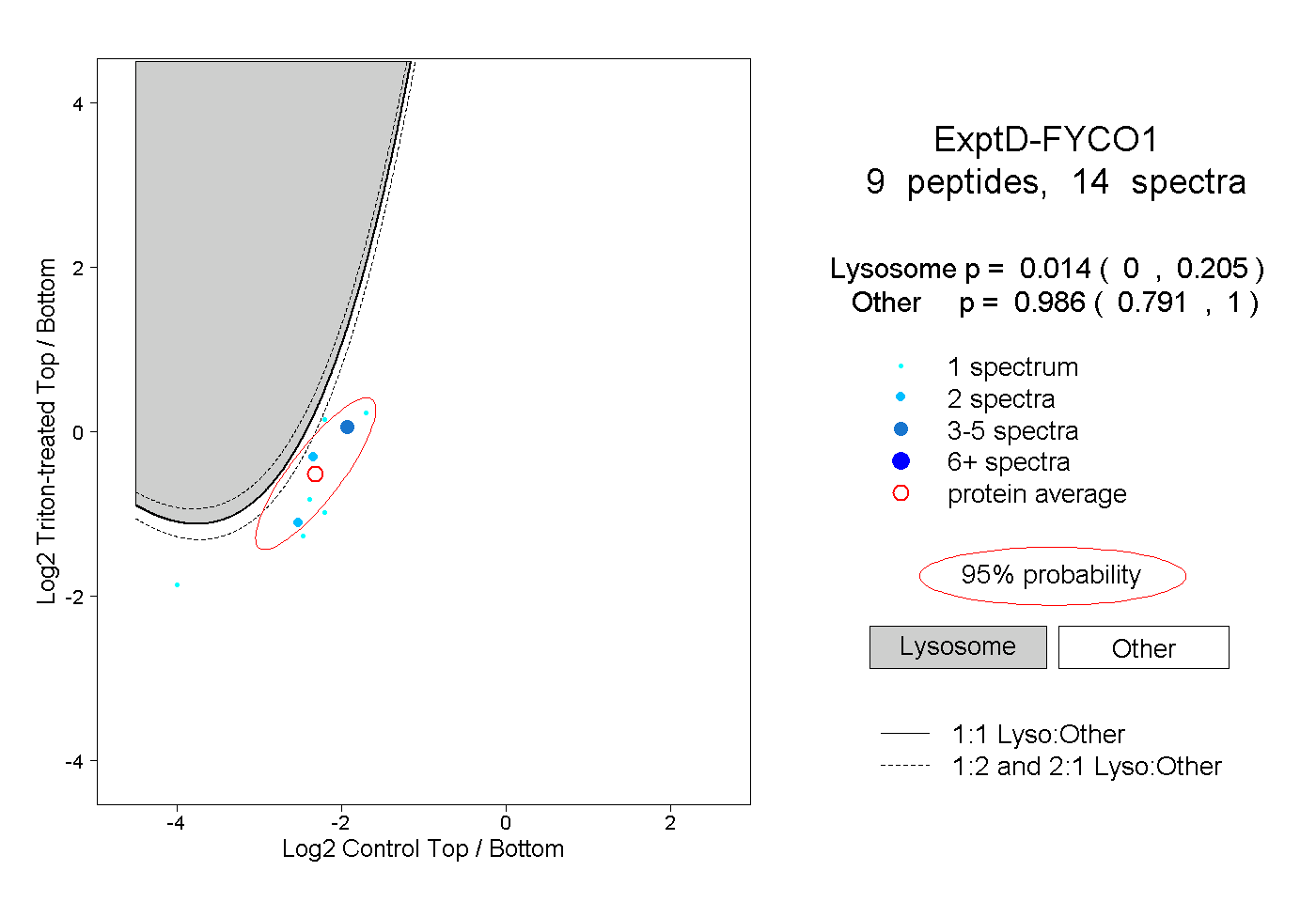 |
0.014 0.000 | 0.205 |
0.986 0.791 | 1.000 |